Location: Chauhan, Legewie, Westermark, Lorenzen and Herzel, 2008 @ 93568094db73 / Chauhan_2008.html
- Author:
- Hanne Nielsen <hnie010@aucklanduni.ac.nz>
- Date:
- 2011-09-02 10:44:46+12:00
- Desc:
- Added details to the HTML file
- Permanent Source URI:
- https://models.cellml.org/workspace/chauhan_legewie_westermark_lorenzen_herzel_2008/rawfile/93568094db734beaae3fb4a43f0ca1e523dcbaf3/Chauhan_2008.html
Model Status
This CellML model will run in OpenCell on the windows machine it was built on to reproduce published results for all graphs but CKI. The model registers errors when run on my mac: does not reproduce published results as there is an error. (Integration failed at t=1.35325, a component of ewt has become <=0) however it reproduces results when run on a PC
Model Structure
The liver regenerates and maintains its function and size after injury by counterbalancing cell death with compensatory cell division. During liver regeneration, injured sites release cytokines, which stimulate normally quiescent hepatocytes to re-enter cell division cycle. Using a mesoscale approach, we have implemented the first mathematical model that describes cytokine-induced dedifferentiation of hepatocytes and the subsequent initiation of DNA synthesis (G0/G1 and G1/S phase transitions of the cell cycle). The model accurately reproduces experimentally measured kinetics of various signaling intermediates and DNA synthesis in hepatocytes for varying degrees of liver damage, in both wild type and knockout backgrounds. Liver regeneration is known to be a robust process, as liver mass reconstitution still occurs in various knockout mice (albeit with different kinetics). We analyze the robustness of the model using methods of control analysis. Moreover, we discuss the system's bandpass filtering properties and delays, which arise from feedbacks and nested feed-forward loops.
The original paper reference is cited below:
'A mesoscale model of G1/S phase transition in liver regeneration.', Anuradha Chauhan, Stefan Legewie, PÃ¥l O. Westermark, Stephan Lorenzenb, Hanspeter Herzel, 2008 Journal of Theoretical Biology, 252, 465-473. PubMed ID: 18339403
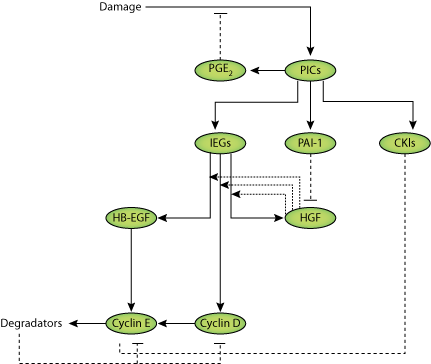
|
| Image illustrating Chauhan 2008 model |

