Butera, Rinzel, Smith, 1999
Model Status
This CellML model represents a single neuron in a network of cells. This cell model is based on "model 1" published in a preceding paper (Butera et al. 1999) which is also described in CellML as a stand alone CellML 1.0 model. To model inter-cellular coupling a synaptic input current Isyn_e has been added to the model.
The MATLAB script (buteraModel.m) should be used to generate a CellML file for multi-cell models. The generated model will import butera_single_cell_1999.cellml repeatedly, once for each of the multiple cells, and it will generate values from a random normal distribution for the required connectivity coefficients between the cells. The main model for the exposure (butera_ten_cell_1999.cellml) was generated from the MATLAB script.
Note: the OpenCell session file for this model may not work when lauched from the model repository. In order to run the session file, you may either clone the workspace or save the session file, and then run the session from your local copy.
Model Structure
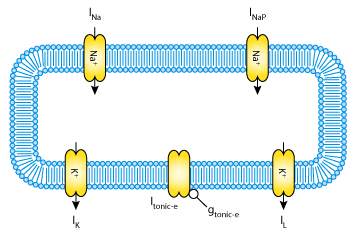 |
| The single cell neuron model is based on a single-compartment Hodgkin-Huxley type formalism. It is composed of five ionic currents across the plasma membrane: a fast sodium current, INa; a delayed rectifier potassium current, IK; a persistent sodium current, INaP; a passive leakage current, IL; and a tonic current, Itonic_e In addition for the multicellular model a synaptic current (Isyn_e) has been added to connect the cells in the network. |
The original paper reference is cited below:
Models of respiratory rhythm generation in the Pre-Botzinger complex. II. Populations Of coupled pacemaker neurons, Butera RJ Jr, Rinzel J, Smith JC, 1999, Journal of Neurophysiology, 82, 398-415. PubMed ID: 10400967

