Schoeberl, Eichler-Jonsson, Gilles, Muller, 2002
Model Status
This CellML model runs in both OpenCell and COR to produce graphs similar to the figures in the published paper. Units have been checked and are consistent. The same model in the BioModels Database was a very helpful guide during the translation process. In order to produce the graphs in the paper, the following rescalings were used: a) concentrations initially in ng/mL were rescaled to molecules per cell (10E-18 x Na/MW with Na = 6.022E23, MW = 6045 and cell volume of 1E-12L) b) first order rate constants were rescaled from /sec to /minute c) second order rate constants were rescaled to /(minute x molecule)
Model Structure
ABSTRACT: We present a computational model that offers an integrated quantitative, dynamic, and topological representation of intracellular signal networks, based on known components of epidermal growth factor (EGF) receptor signal pathways. The model provides insight into signal-response relationships between the binding of EGF to its receptor at the cell surface and the activation of downstream proteins in the signaling cascade. It shows that EGF-induced responses are remarkably stable over a 100-fold range of ligand concentration and that the critical parameter in determining signal efficacy is the initial velocity of receptor activation. The predictions of the model agree well with experimental analysis of the effect of EGF on two downstream responses, phosphorylation of ERK-1/2 and expression of the target gene, c-fos.
The original paper reference is cited below:
Computational modeling of the dynamics of the MAP kinase cascade activated by surface and internalized EGF receptors, Birgit Schoeberl, Claudia Eichler-Jonsson, Ernst Dieter Gilles, Gertraud Muller, 2002, Nature Biotechnology, 20, 370 - 375. PubMed ID: 11923843
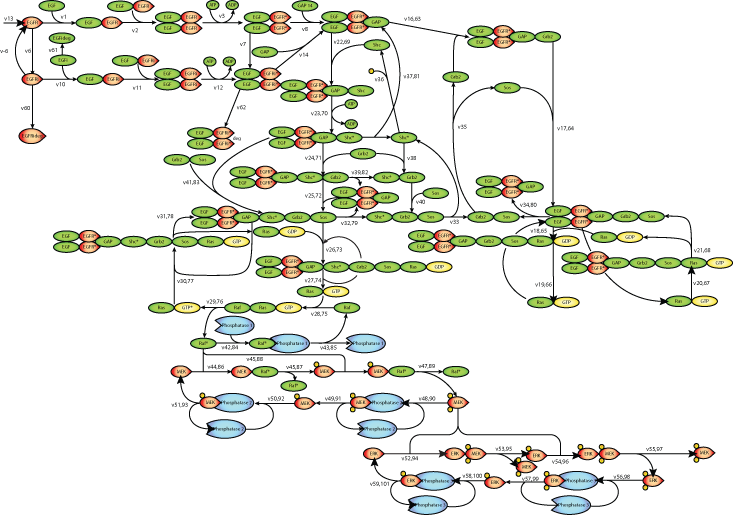 |
| Scheme of the EGF receptor-induced MAP kinase cascade. |

