Palecek, Horwitz, Lauffenburger, 1999
Model Status
This model contains partial differentials, which do not fit within the CellML 1.0 or 1.1 specifications.
Please note, because the parameters for the attachment and detachment phases of the model are given the same identifiers, the model in the CellML format has been divided into two separate models, one for attachment and the other for the detachment phase. This particular version of the model is for the detachment phase.
Model Structure
The process of cell migration is a spatially and temporally coordinated process of events. Under many different conditions, the rate limiting event of cell migration is the rate at which the cell and the substratum detach at the cell rear.
In order to invetigate this cellular process further, Palecek et al. have devloped a mathematical model to integrate how the biophysical and biochemical interactions between integrins, the cytoskelton, and the extracellular matrix (ECM) affect rear retraction and linkage dissociation mechanisms (see the figure below). The model also examines how applied forces and integrin clustering affect retracion kinetics, and it predicts two distinct phenotypes. In the first, detachment is rapid, dominated by integrin-ECM dissociation, and it occurs at high forces or low adhesiveness. In the second phenotype, detachment is slower, and it occurs at low forces or high adhesiveness. The model also helps to explain why some cell types, such as leukocytes or keratocytes, are able to detach easily and move quickly while other cell types, such as fibroblasts, migrate more slowly and release more integrins during detachment.
Because the parameters for the attachment and detachment phases of the model are given the same identifiers, the model in the CellML format has been divided into two separate models, one for attachment and the other for the detachment phase.
The complete original paper reference is cited below:
Kinetic Model for Integrin-mediated Adhesion Release During Cell Migration, Sean P. Palecek, Alan F. Horwitz, and Douglas A. Lauffenburger, 1999, Annals of Biomedical Engineering , 27, 219-235. (A PDF version of the article is available to subscribers on the Annals of Biomedical Engineering website.) PubMed ID: 10199699
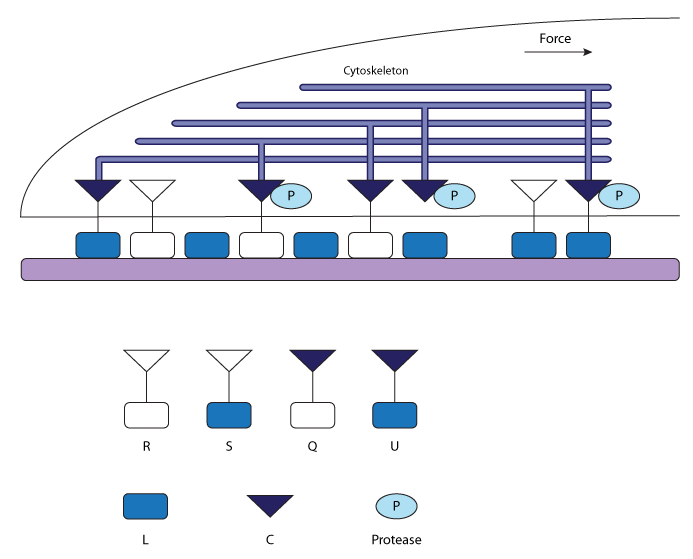 |
| Schematic representation of linkages between the cell and the substratum during rear reaction. Integrins can exist in four states: unbound (R); bound to the ECM ligand (S); bound to the cytoskeleton (Q); or bound to both the ECM and the cytoskelton (U). Ligands can either be free (L) or bound to integrins (S). Cytoskeletal elements can also be free (C) or bound to integrins (Q). Upon activation, protease (P) can cleave integrin-cytoskeletal linkages. |

