Wu, Jeneson, Beard, 2006
Model Status
This CellML version of the model runs in OpenCell and COR to reproduce the original matlab code, which it has been based off. The units are consistent throughout.
Model Structure
Abstract: Data from 31P-nuclear magnetic resonance spectroscopy of human forearm flexor muscle were analyzed based on a previously developed model of mitochondrial oxidative phosphorylation [PLoS Comp. Bio. (2005) 1:e36] to test the hypothesis that substrate level (concentrations of ADP and inorganic phosphate) represents the primary signal governing the rate of mitochondrial ATP synthesis and maintaining the cellular ATP hydrolysis potential in skeletal muscle. Model-based predictions of cytoplasmic concentra-tions of phosphate metabolites (ATP, ADP, and inorganic phosphate) matched data obtained from 20 healthy volunteers and indicated that as work rate is varied from rest to maximal exercise commensurate increases in the rate of mitochondrial ATP synthesis are effected by changes in concentrations of avail-able ADP and inorganic phosphate. Additional data from patients with a defect of complex I of the res-piratory chain and a patient with a deficiency in the mitochondrial adenine nucleoside translocase were also predicted the by the model by making the appropriate adjustments to the activities of the affected proteins associates with the defects, providing both further validation of the biophysical model of the control of oxidative phosphorylation and insight into the impact of these diseases on the ability of the cell to maintain its energetic state.
The complete original paper reference is cited below:
Oxidative ATP Synthesis in Skeletal Muscle is Controlled by Substrate Feedback, Fan Wu, Jeroen A.L. Jeneson, Daniel A. Beard, 2006, American Journal of Physiology - Cell Physiology, 292, C115-24 PubMed ID: 16837647
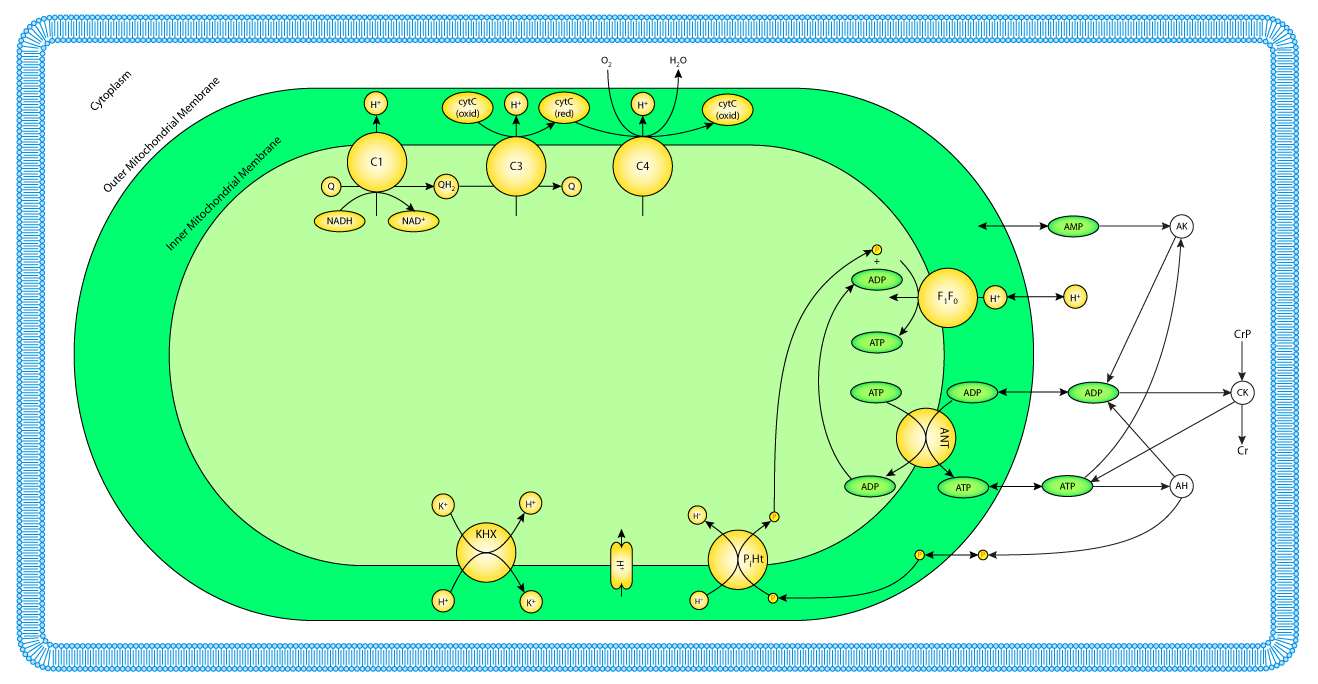 |
| Schematic diagram of the components included in the computational model of oxidative phosphorylation in skeletal muscular cells. All reactions and mass transport take place in three compartments: cytoplasm, mitochondrial intermembrane space, and the mitochondrial matrix. |

