Obeyesekere, Zimmerman, Tecarro, Auchmuty, 1999
Model Status
This CellML version of the model has been checked in COR and PCEnv and the model runs to replicate the original published results as depicted in figure 3a of the paper. The units have been checked and are consistent. Additional variables 'free-E2F' and 'unphos-RB' were added for plotting.
Model Structure
ABSTRACT: A modified version of a previously developed mathematical model [Obeyesekere et al., Cell Prolif. (1997)] of the G1-phase of the cell cycle is presented. This model describes the regulation of the G1-phase that includes the interactions of the nuclear proteins, RB, cyclin E, cyclin D, cdk2, cdk4 and E2F. The effects of the growth factors on cyclin D synthesis under saturated or unsaturated growth factor conditions are investigated based on this model. The solutions to this model (a system of nonlinear ordinary differential equations) are discussed with respect to existing experiments. Predictions based on mathematical analysis of this model are presented. In particular, results are presented on the existence of two stablesolutions, i.e., bistability within the G1-phase. It is shown that this bistability exists under unsaturated growth factor concentration levels. This phenomenon is very noticeable if the efficiency of the signal transduction, initiated by the growth factors leading to cyclin D synthesis, is low. The biological significance of this result as well as possible experimental designs to test these predictions are presented.
The original paper reference is cited below:
A model of cell cycle behavior dominated by kinetics of a pathway stimulated by growth factors, Mandri N. Obeyesekere, Stuart O. Zimmerman, Edwin S. Tecarro and Giles Auchmuty, 1999, Bulletin of Mathematical Biology, 61, 917-934. PubMed ID: 17886749
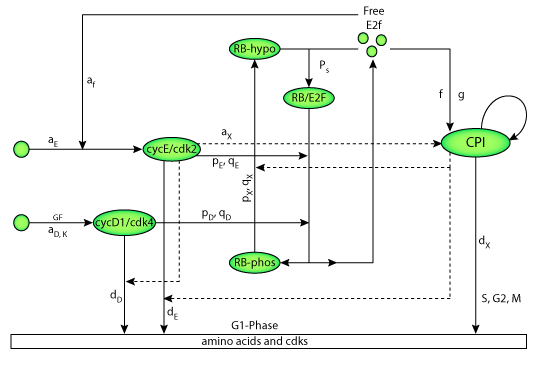 |
| Flowchart illustrating the protein interactions described by the model. |

