Ciliberto, Novak, Tyson, 2003
Model Status
This CellML version of the model has been checked in COR and OpenCell and the model runs to replicate the results in the original published paper. The model was checked against the XPP code as there is an inconsistency between the code and the published equation for the variable 'PSwe1'. The units have been checked and they are consistent.
Model Structure
ABSTRACT: The morphogenesis checkpoint in budding yeast delays progression through the cell cycle in response to stimuli that prevent bud formation. Central to the checkpoint mechanism is Swe1 kinase: normally inactive, its activation halts cell cycle progression in G2. We propose a molecular network for Swe1 control, based on published observations of budding yeast and analogous control signals in fission yeast. The proposed Swe1 network is merged with a model of cyclin-dependent kinase regulation, converted into a set of differential equations and studied by numerical simulation. The simulations accurately reproduce the phenotypes of a dozen checkpoint mutants. Among other predictions, the model attributes a new role to Hsl1, a kinase known to play a role in Swe1 degradation: Hsl1 must also be indirectly responsible for potent inhibition of Swe1 activity. The model supports the idea that the morphogenesis checkpoint, like other checkpoints, raises the cell size threshold for progression from one phase of the cell cycle to the next.
The original paper reference is cited below:
Mathematical model of the morphogenesis checkpoint in budding yeast, Andrea Ciliberto, Bela Novak, and John J. Tyson, 2003, The Journal of Cell Biology , 163, 1243-1254. PubMed ID: 14691135
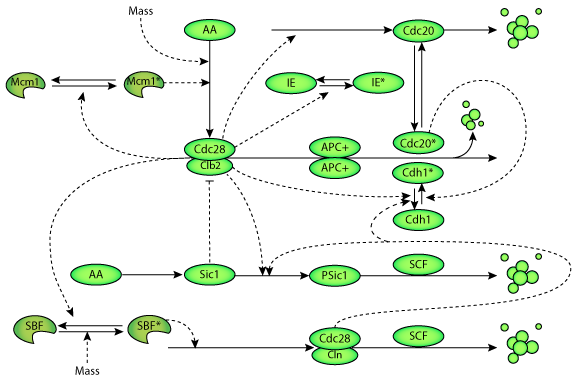 |
| A schematic diagram of the molecular mechanisms underlying the regulation of the cell cycle in budding yeast. |
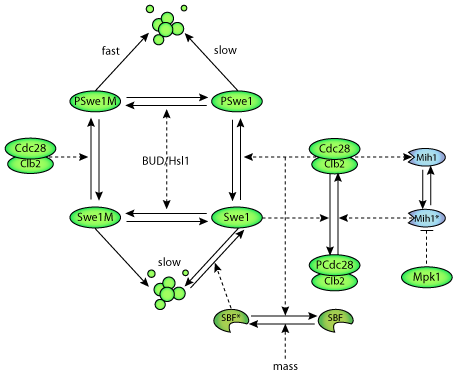 |
| A schematic diagram of the Swe1 box - a process which is central to the morphogenesis checkpoint in budding yeast. |

