Location: Metabolic Component Library @ af10084679bb / components.html
- Author:
- Matthias K?nig <matthias.koenig@charite.de>
- Date:
- 2013-04-09 15:08:59+02:00
- Desc:
- Documentation of Reaction Mechanism. Images for basic mechanisms created.
- Permanent Source URI:
- https://models.cellml.org/w/matthiaskoenig/MetabolicComponentLibrary/rawfile/af10084679bb928d763a8ff441ed93d41aa59126/components.html
Metabolic Component Library for CellML
Matthias Koenig [1] & Randall Britten [2][1] Charite Berlin, matthias.koenig@charite.de
[2] Auckland Bioengineering Institute
Implementation of standardized reaction kinetics in CellML for reuse in metabolic models. The library covers the metabolic function definitions for processes from COPASI 4.8 (Bulid 35).
Naming Conventions
Substrate is named S (S1, S2, ... in case of multiple substrates)Products is named P (P1, P2, ... in case of multiple productes)
Inhibitor is named I (I1, I2, ...)
Activator is named A (A1, A2, ...)
Modifier is named M (M1, M2, ...)
Km values are subscripted with the respective substrates or products (P -> Km_P, S -> Km_S, P1 -> Km_P1, ...)
Analog convention for inhibition constants Ki and activation constants Ka.
Units
Time: secondVolume Unit: liter
Quantity Unit: mmol
Components
AllostericInhibitionEmpirical
CellML
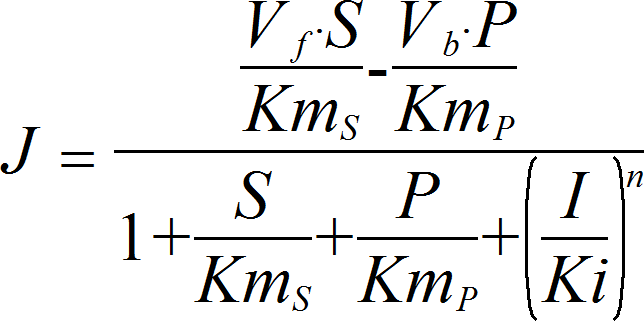
UniUniRev_I
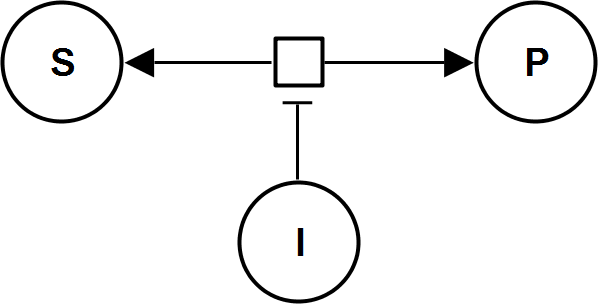
AllostericInhibitionMWC
CellML
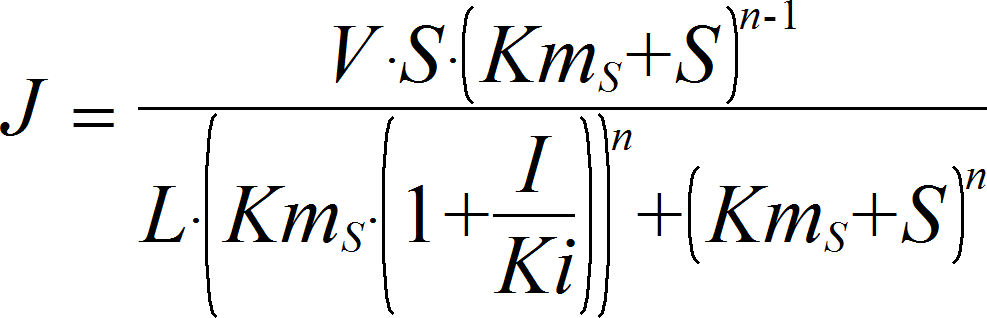
UniUniIrrev_I
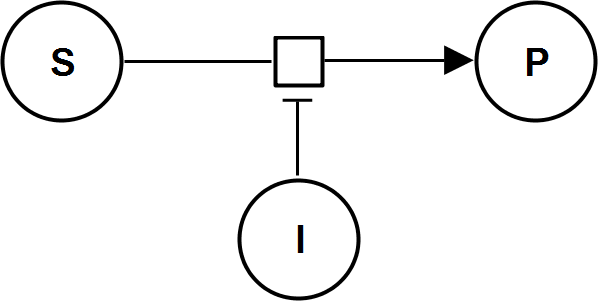 UniBiIrrev_I
UniBiIrrev_I
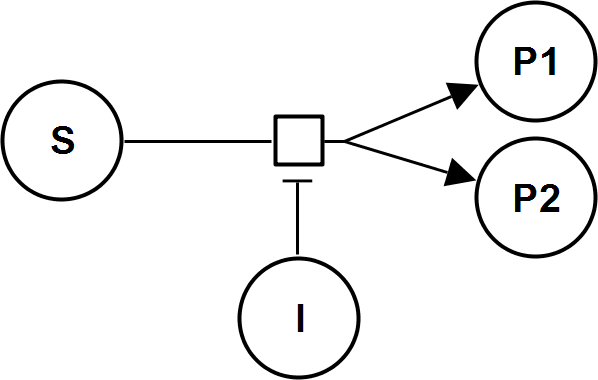 UniNullIrrev_I
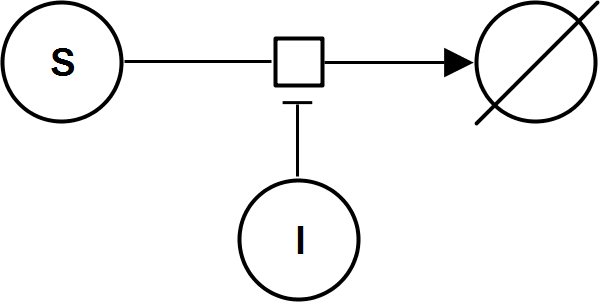
UniNullIrrev_I
BiIrrev
CellML
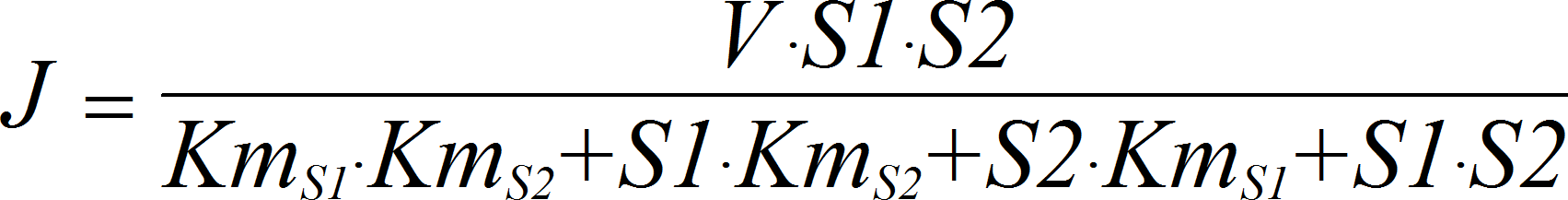
BiNullIrrev
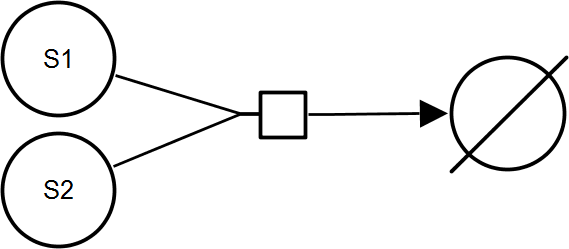 BiBiIrrev
BiBiIrrev
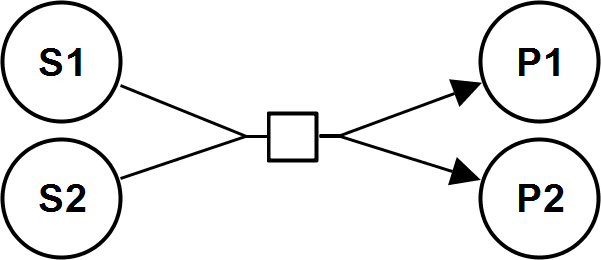 BiUniIrrev
BiUniIrrev
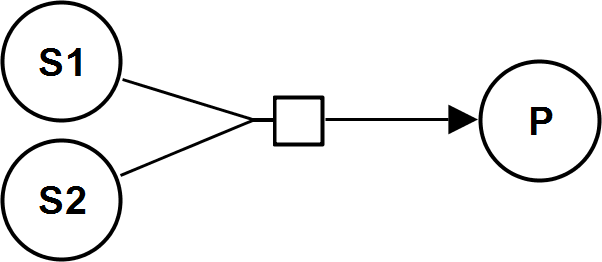
CatalyticActivationIrrev
CellML
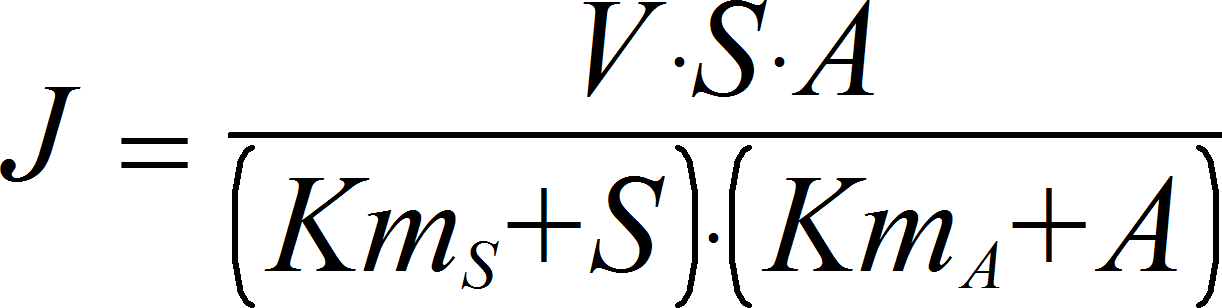
UniUniIrrev_A
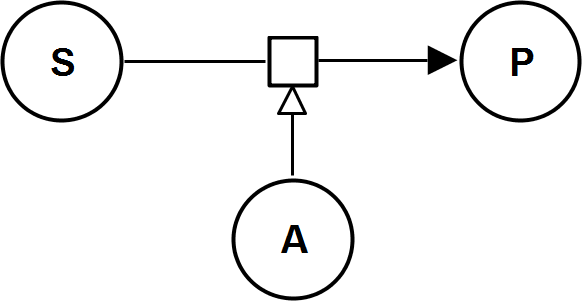 UniBiIrrev_A
UniBiIrrev_A
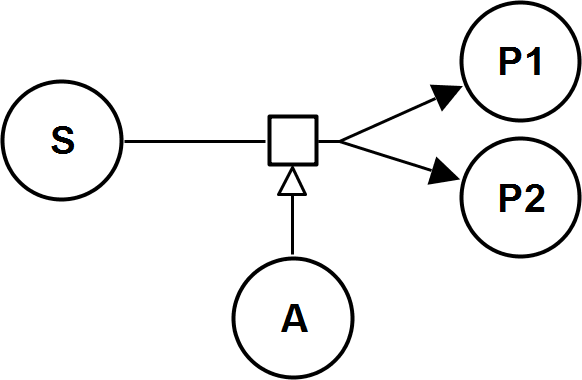 UniNullIrrev_A
UniNullIrrev_A
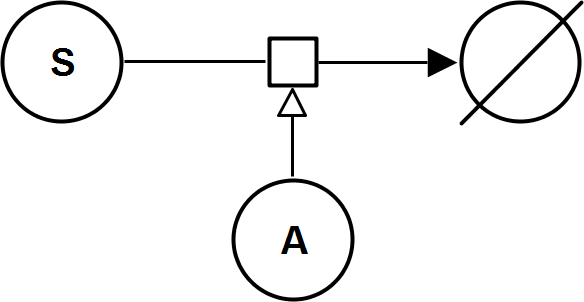
CatalyticActivationRev
CellML
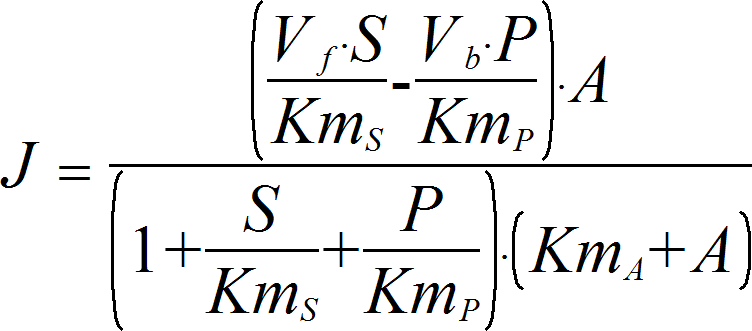
UniUniRev_A
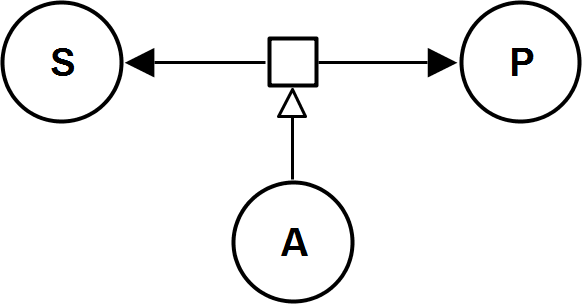
CompetitiveInhibitionIrrev
CellML
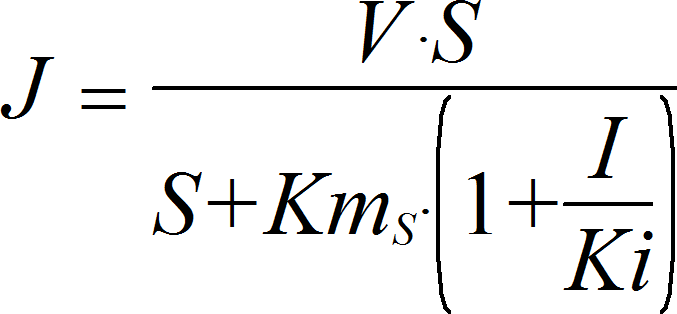
UniUniIrrev_I
 UniBiIrrev_I
UniBiIrrev_I
 UniNullIrrev_I
UniNullIrrev_I

CompetitiveInhibitionRev
CellML
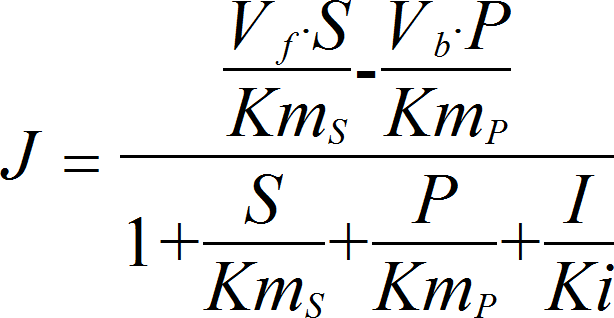
UniUniRev_I

ConstantFluxIrrev
CellML
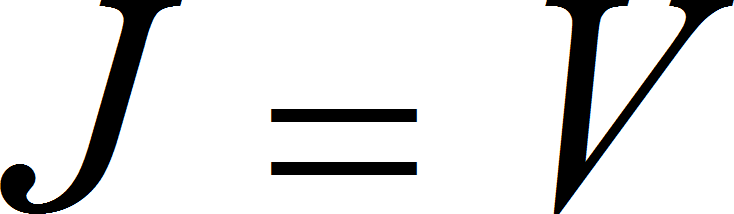
UniNullIrrev
 UniUniIrrev
UniUniIrrev
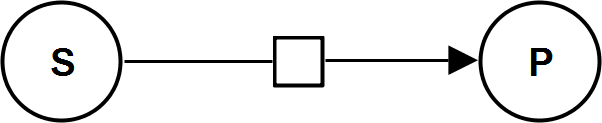 UniBiIrrev
UniBiIrrev
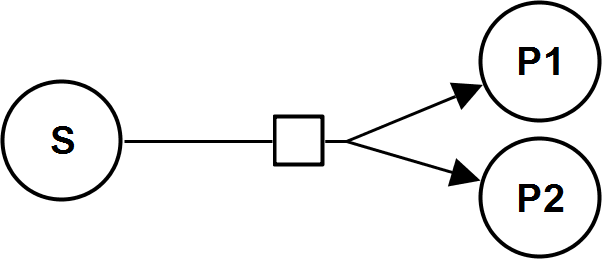 BiBiIrrev
BiBiIrrev
 BiUniIrrev
BiUniIrrev

Hill1ModifierRev
CellML
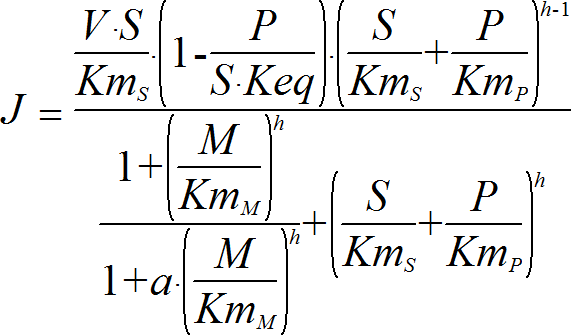
UniUniRev_M
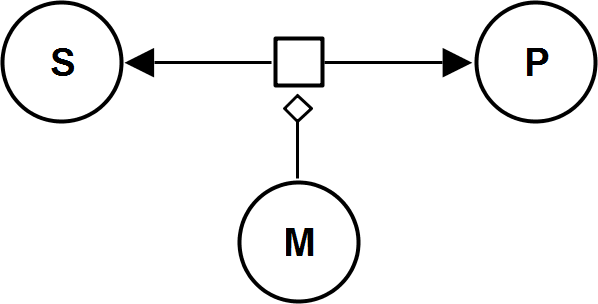
Hill2ModifierRev
CellML
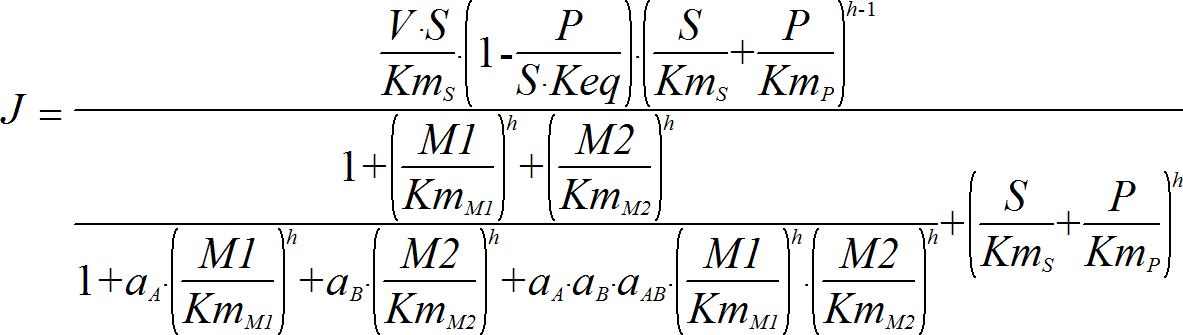
UniUniRev_M2
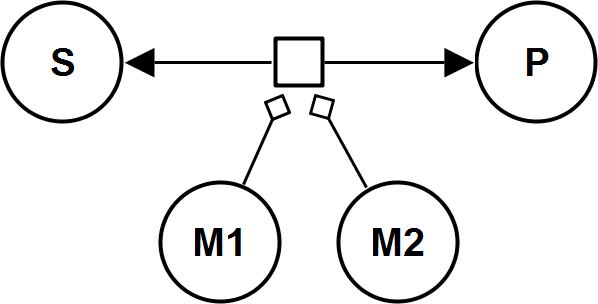
HillIrrev
CellML
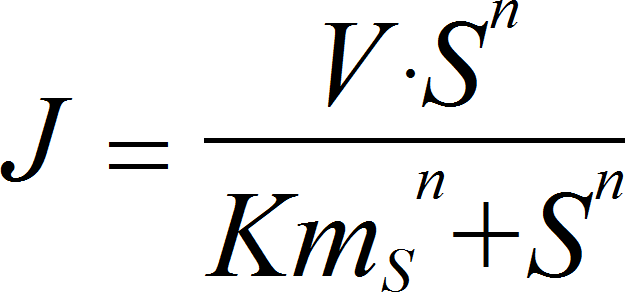
UniNullIrrev
 UniUniIrrev
UniUniIrrev
 UniBiIrrev
UniBiIrrev

HillRev
CellML
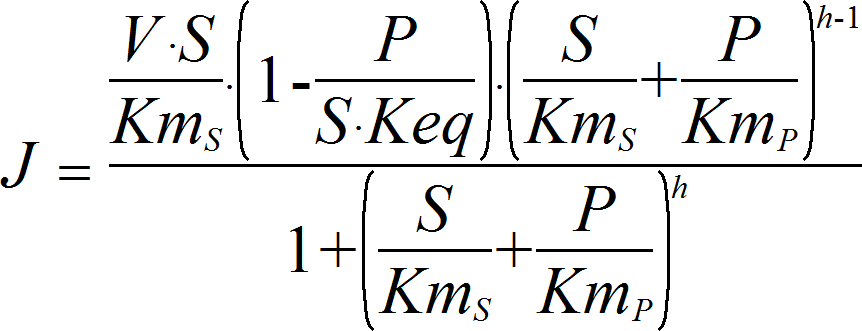
UniUniRev
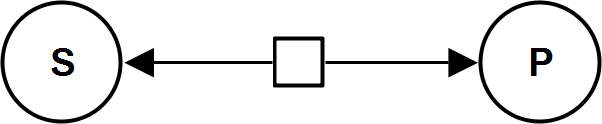
HyperbolicModifierIrrev
CellML
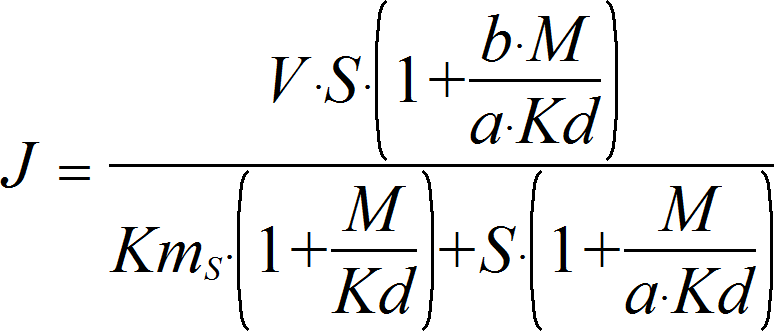
UniNullIrrev_M
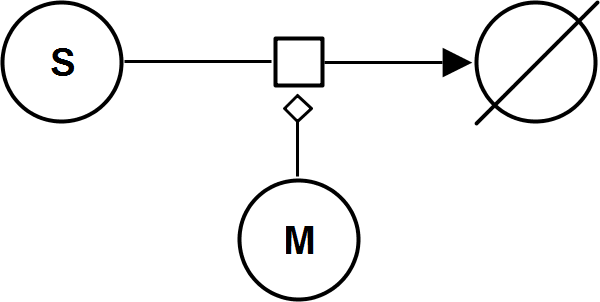 UniUniIrrev_M
UniUniIrrev_M
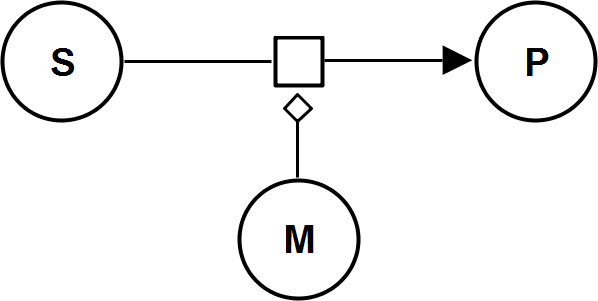 UniBiIrrev_M
UniBiIrrev_M
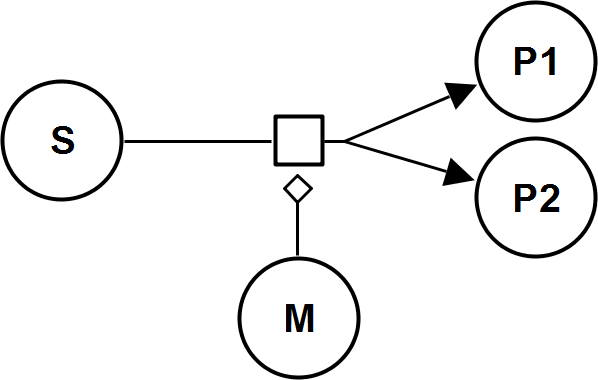
HyperbolicModifierRev
CellML
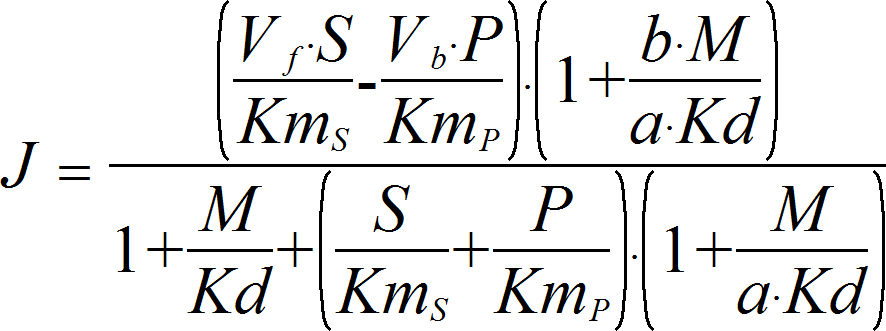
UniUniRev_M

IsoUniUni
CellML
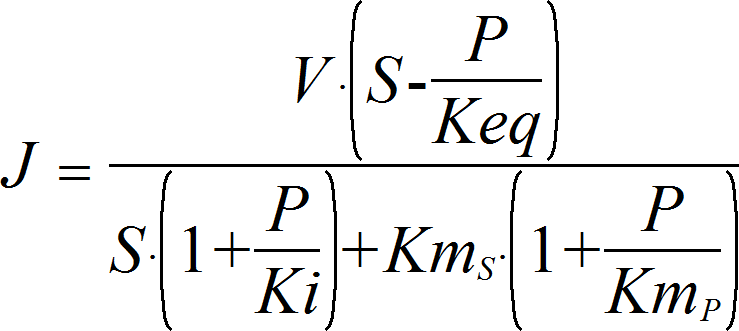
UniUniRev

MassActionBiBiRev
CellML
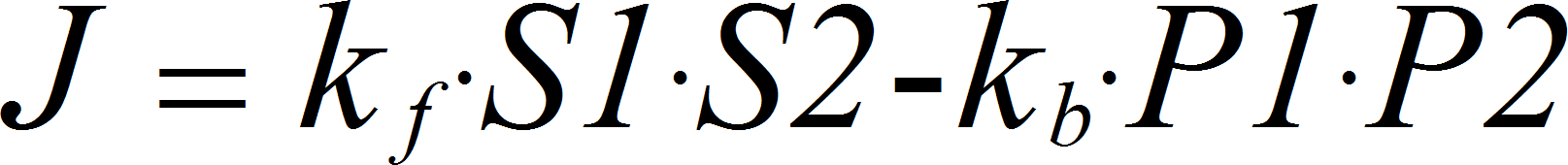
BiBiRev
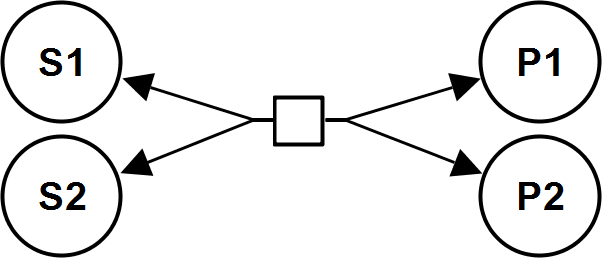
MassActionBiIrrev
CellML
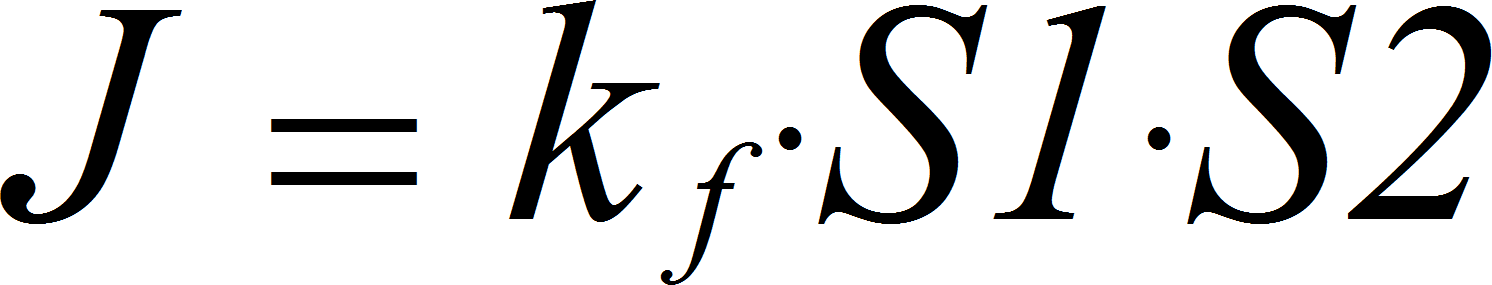
BiNullIrrev
 BiBiIrrev
BiBiIrrev
 BiUniIrrev
BiUniIrrev

MassActionBiUniRev
CellML
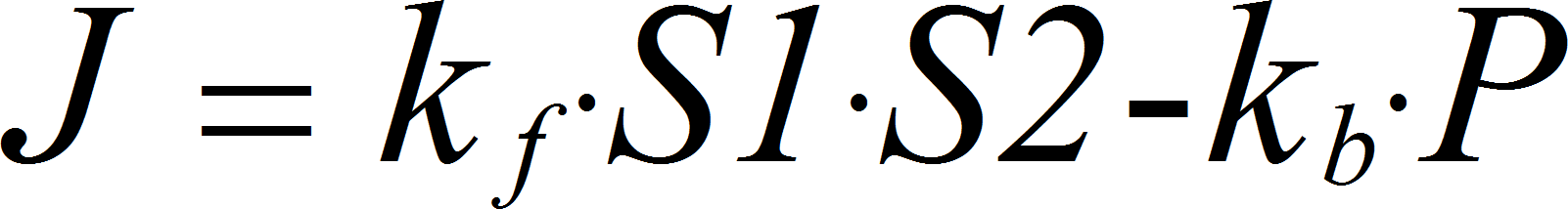
BiUniRev
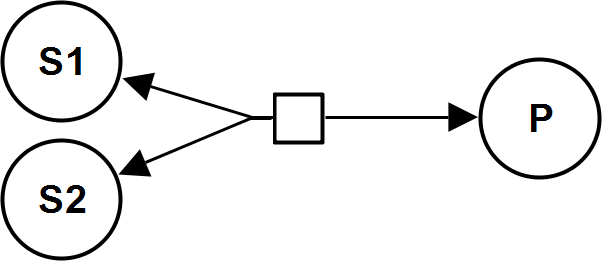
MassActionUniBiRev
CellML
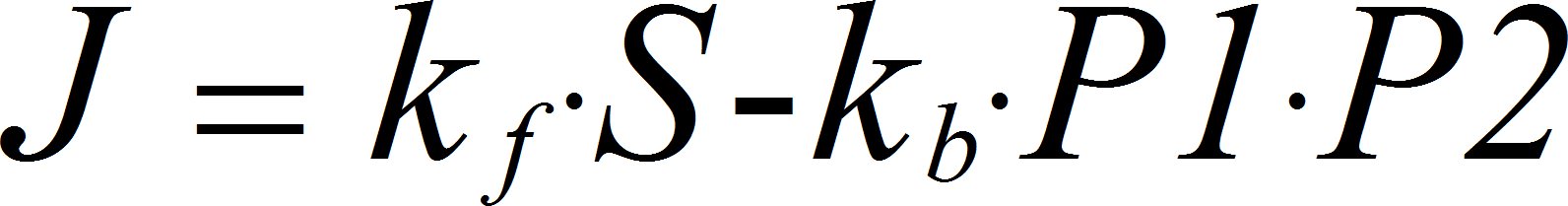
UniBiRev
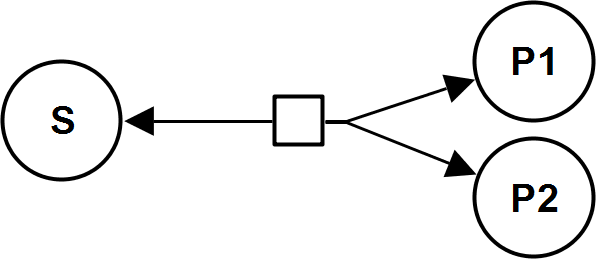
MassActionUniIrrev
CellML
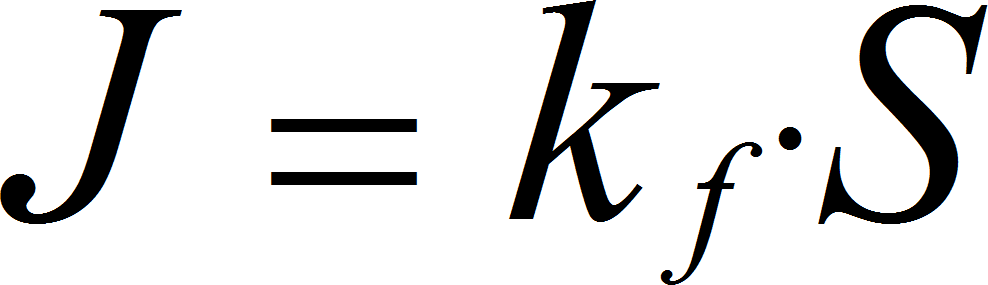
UniNullIrrev
 UniUniIrrev
UniUniIrrev
 UniBiIrrev
UniBiIrrev

MassActionUniUniRev
CellML
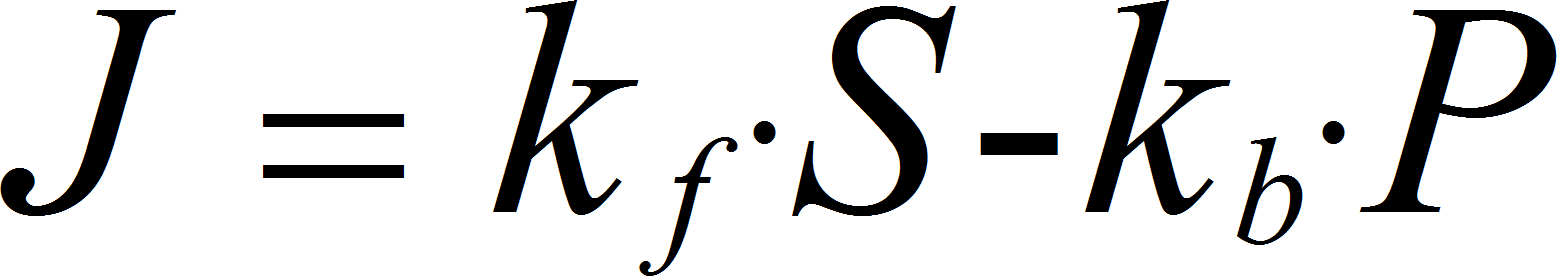
UniUniRev

MichaelisMentenIrrev
CellML
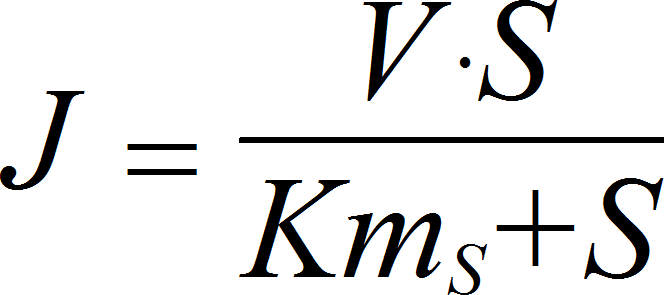
UniNullIrrev
 UniUniIrrev
UniUniIrrev
 UniBiIrrev
UniBiIrrev

MichaelisMentenRev
CellML
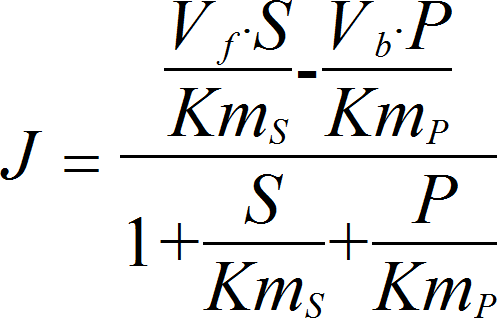
UniUniRev

MixedActivationIrrev
CellML
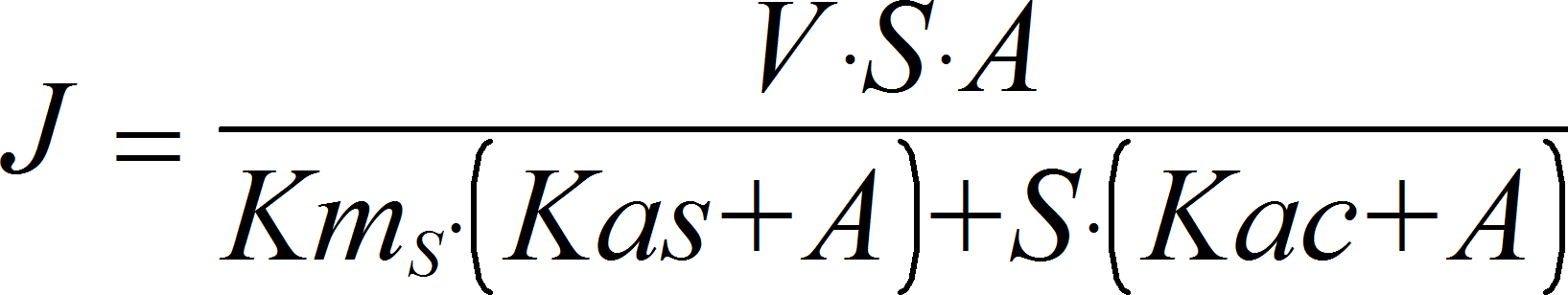
UniUniIrrev_A
 UniBiIrrev_A
UniBiIrrev_A
 UniNullIrrev_A
UniNullIrrev_A

MixedActivationRev
CellML
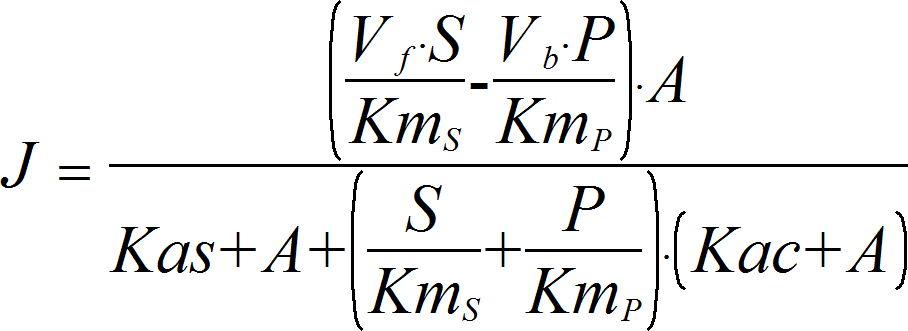
UniUniRev_A

MixedInhibitionIrrev
CellML
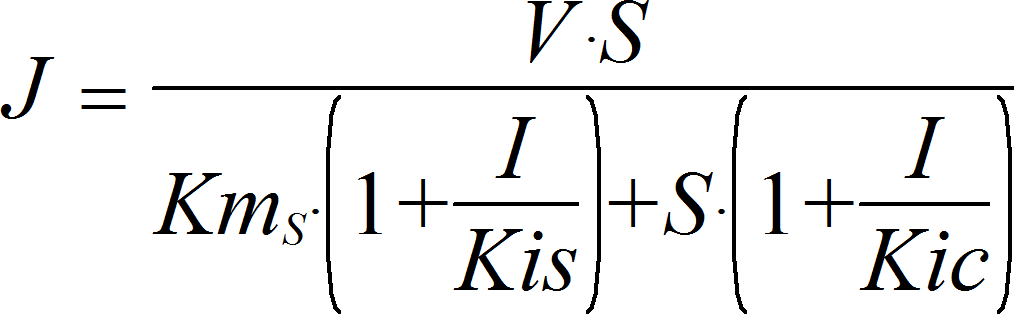
UniUniIrrev_I
 UniBiIrrev_I
UniBiIrrev_I
 UniNullIrrev_I
UniNullIrrev_I

MixedInhibitionRev
CellML
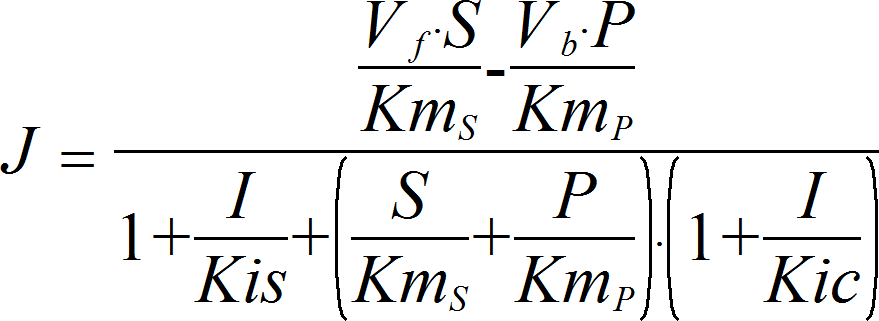
UniUniRev_I

NoncompetitiveInhibitionIrrev
CellML
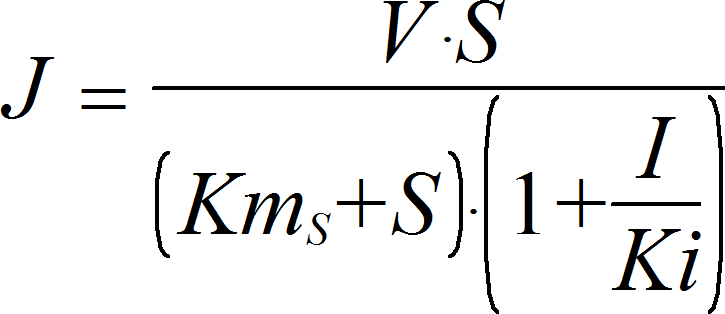
UniUniIrrev_I
 UniBiIrrev_I
UniBiIrrev_I
 UniNullIrrev_I
UniNullIrrev_I

NoncompetitiveInhibitionRev
CellML
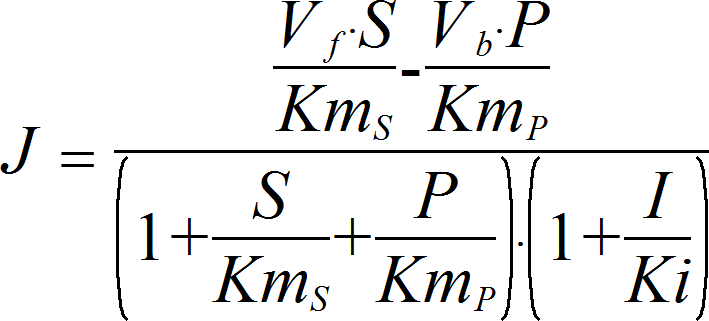
UniUniRev_I

OrderedBiBi
CellML
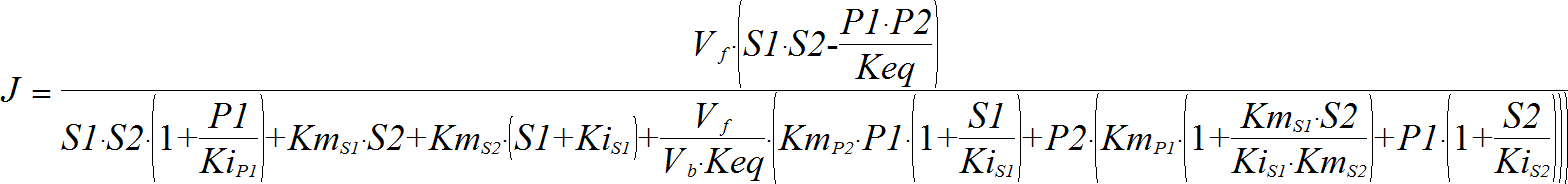
BiBiRev

OrderedBiUni
CellML
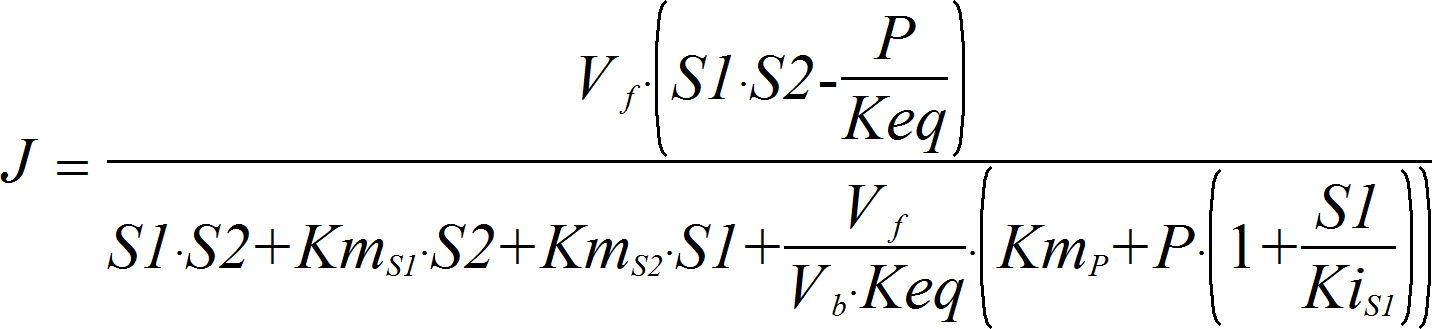
BiUniRev

OrderedUniBi
CellML
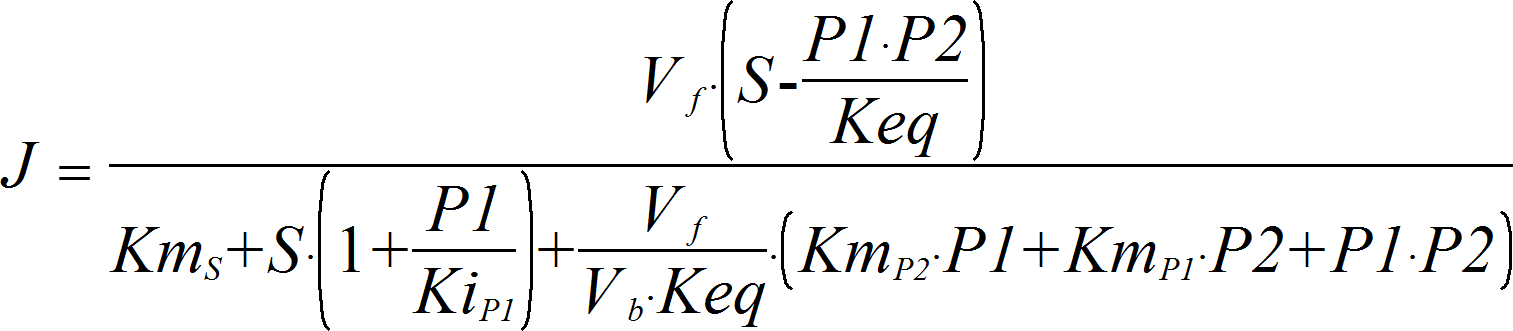
UniBiRev

PingPongBiBi
CellML
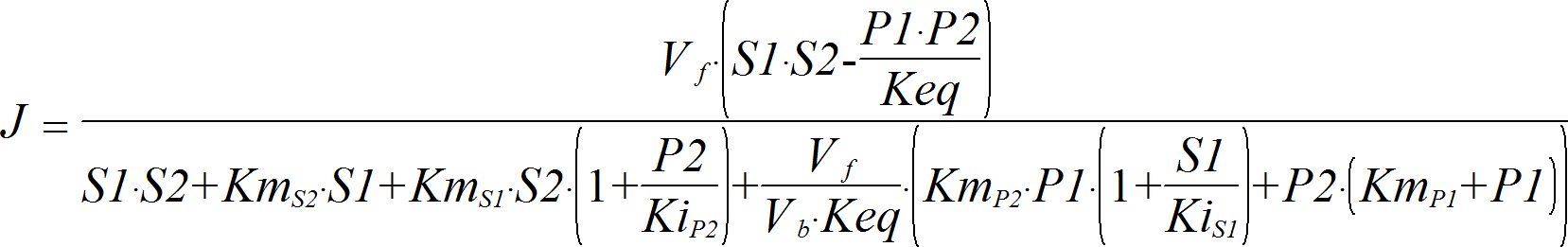
BiBiRev

SpecificActivationIrrev
CellML
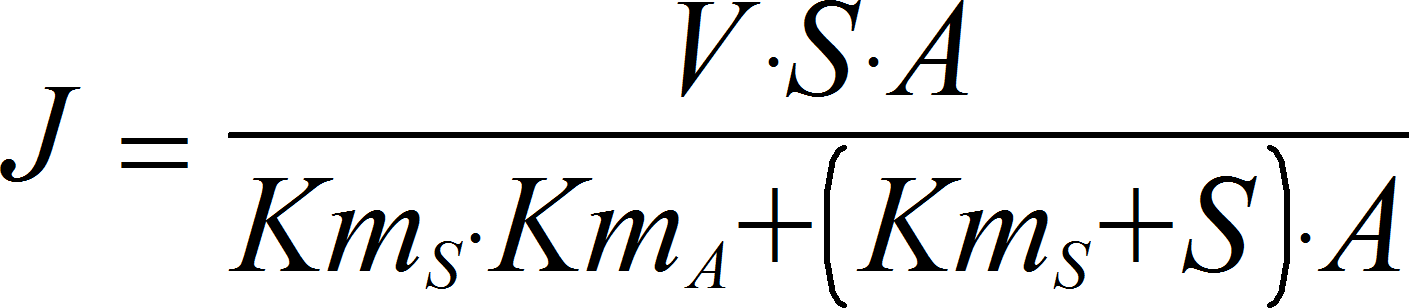
UniUniIrrev_A
 UniBiIrrev_A
UniBiIrrev_A
 UniNullIrrev_A
UniNullIrrev_A

SpecificActivationRev
CellML
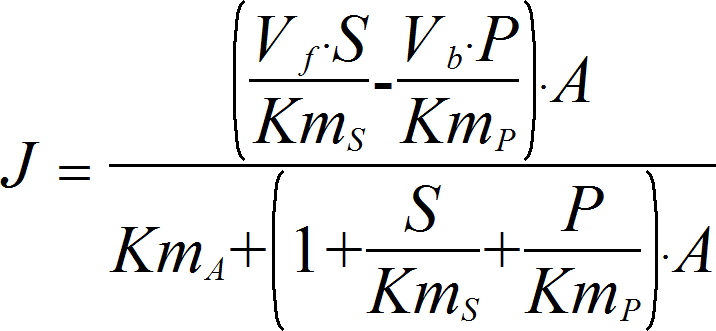
UniUniRev_A

SubstrateActivationIrrev
CellML
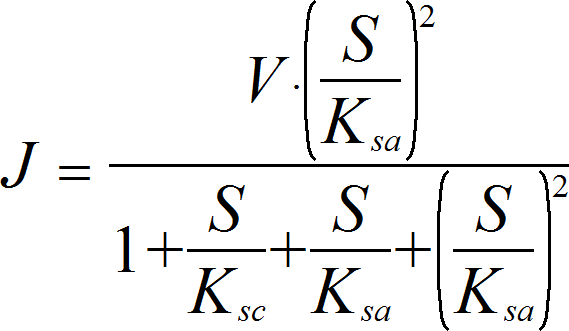
UniNullIrrev
 UniUniIrrev
UniUniIrrev
 UniBiIrrev
UniBiIrrev

SubstrateInhibitionIrrev
CellML
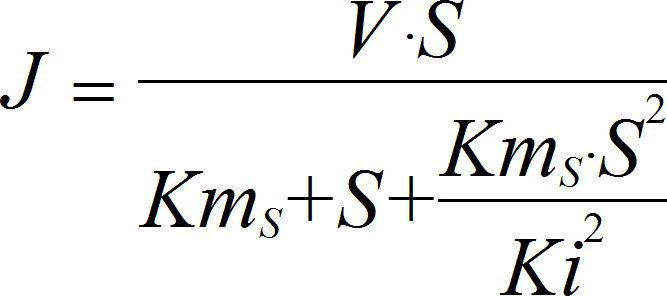
UniNullIrrev
 UniUniIrrev
UniUniIrrev
 UniBiIrrev
UniBiIrrev

SubstrateInhibitionRev
CellML
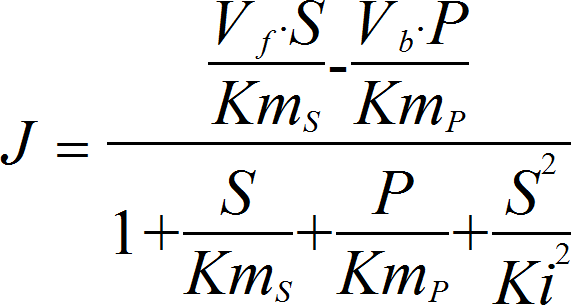
UniUniRev

UncompetitiveInhibitionIrrev
CellML
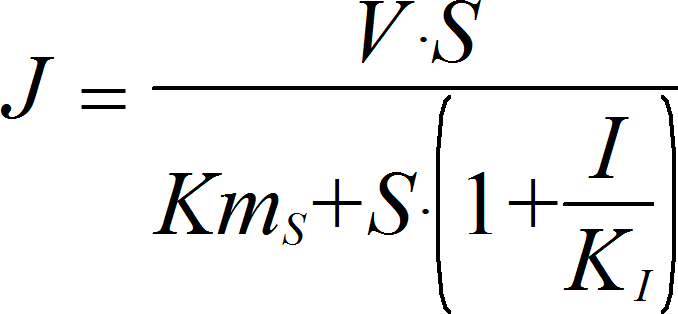
UniUniIrrev_I
 UniBiIrrev_I
UniBiIrrev_I
 UniNullIrrev_I
UniNullIrrev_I


